About me
I am a postdoctoral researcher in the Weisberg and LeBoldus labs in the Botany and Plant Pathology department at Oregon State University.
My research focus on comparative genomics analyses of fungal plant pathogens. Using bioinformatics techniques, I am interested in exploring genome dynamics of fungal plant pathogens to uncover mechanisms of adapation and pahogenicity. In particular, I try to answer questions related to “how pathogens evolve to overcome host resistance?”
In my current research, I use comparative genomics of to investigate how strains of the Eastern Filbert Blight pathogen Anisogramma anomala are capable to overcome ‘Gasaway’ resistance in hazelnut cultivars.
My academic background
My academic journey is quite unique. I earned a BS in Computer Science from the State University of Mato Grosso do Sul, Brazil. Following this, I completed my MS in Computer Science at the Federal University of Mato Grosso do Sul, Brazil. During my master’s, I delved into computational complexity and combinatorial optimization, focusing on a theoretical problem called the maximum similarity partitioning problem. I demonstrated that this problem is NP-complete in the strong sense and proposed greedy heuristics to address it.
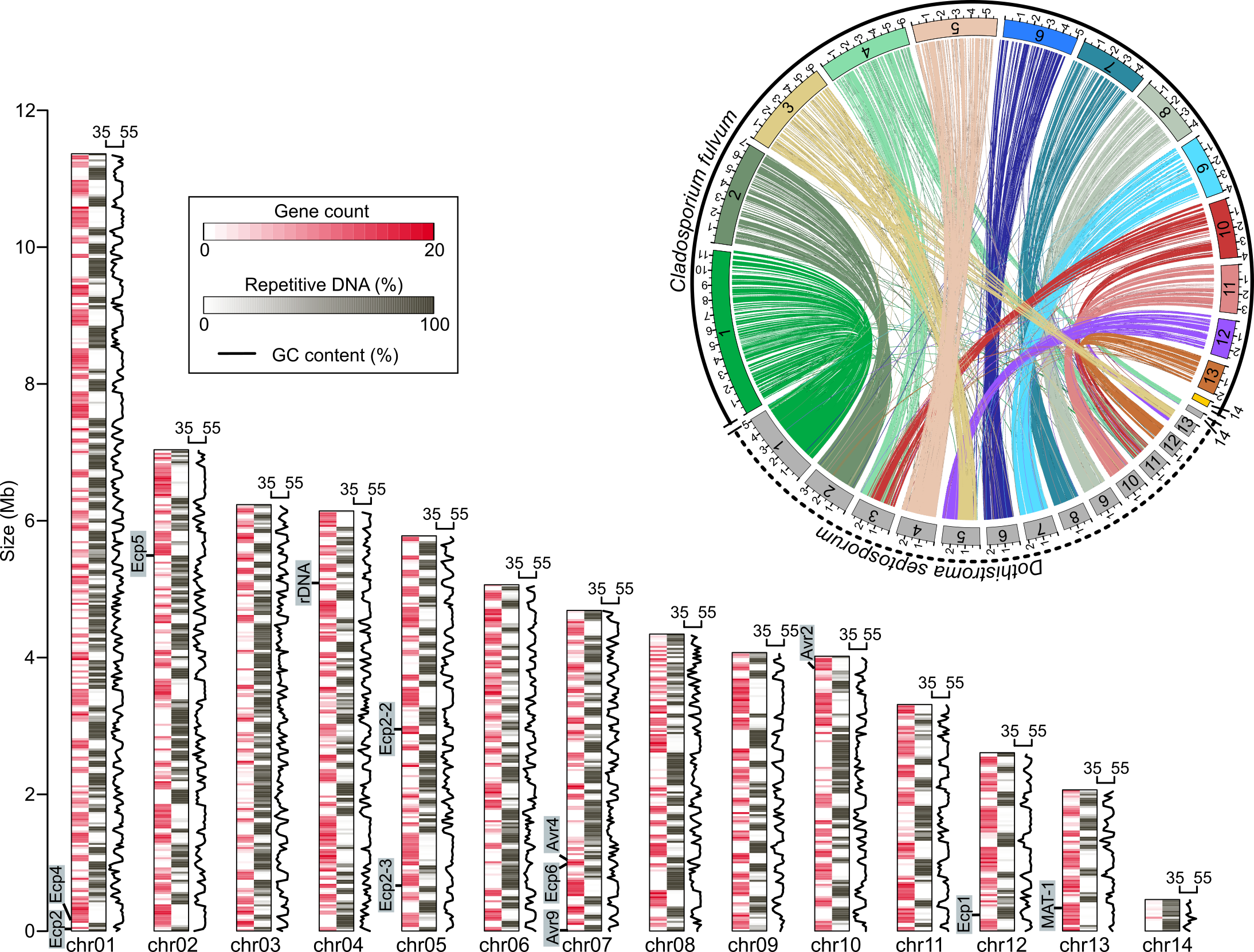
For my PhD, I took a drastic turn. I earned my PhD in Integrative Genetics and Genomics at UC Davis, working in the Stergiopoulos lab. My doctoral research centered on investigating the genome organization and structural variations that influence the evolution of fungal pathogens, specifically the tomato pathogen Cladosporium fulvum and the grape powdery mildew pathogen Erysiphe necator.
To assist in my research, I use a great variety of computational tools. Often, I integrate these tools to build bioinformatics workflows based on Snakemake and bioconda.

